_73.png)
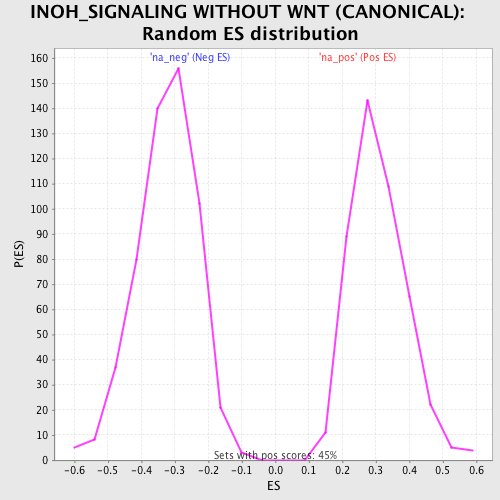
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | INOH_SIGNALING WITHOUT WNT (CANONICAL) |
| Enrichment Score (ES) | -0.70352876 |
| Normalized Enrichment Score (NES) | -2.173785 |
| Nominal p-value | 0.0 |
| FDR q-value | 2.3085259E-4 |
| FWER p-Value | 0.0040 |
_73.png)
| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CUL1 | 1539 | 5.507 | -0.0395 | No | ||
| 2 | DVL3 | 4485 | 2.919 | -0.1576 | No | ||
| 3 | TCF3 | 4510 | 2.909 | -0.1425 | No | ||
| 4 | YWHAG | 5501 | 2.556 | -0.1734 | No | ||
| 5 | PYGO1 | 7469 | 1.995 | -0.2520 | No | ||
| 6 | CTNNBIP1 | 13306 | 0.391 | -0.5161 | No | ||
| 7 | SFN | 14174 | -0.015 | -0.5556 | No | ||
| 8 | FRAT2 | 14394 | -0.152 | -0.5647 | No | ||
| 9 | TLE6 | 15459 | -0.851 | -0.6085 | No | ||
| 10 | CTNNB1 | 16702 | -1.841 | -0.6549 | No | ||
| 11 | TLE1 | 17016 | -2.145 | -0.6572 | No | ||
| 12 | TCF7L2 | 18032 | -3.133 | -0.6861 | Yes | ||
| 13 | DVL1 | 18290 | -3.417 | -0.6787 | Yes | ||
| 14 | AXIN2 | 18561 | -3.716 | -0.6703 | Yes | ||
| 15 | YWHAB | 18690 | -3.865 | -0.6546 | Yes | ||
| 16 | TLE3 | 19490 | -5.019 | -0.6631 | Yes | ||
| 17 | AXIN1 | 19618 | -5.227 | -0.6398 | Yes | ||
| 18 | DVL2 | 19696 | -5.330 | -0.6136 | Yes | ||
| 19 | APC | 19779 | -5.445 | -0.5869 | Yes | ||
| 20 | TLE2 | 20244 | -6.380 | -0.5726 | Yes | ||
| 21 | TCF7 | 20577 | -7.144 | -0.5479 | Yes | ||
| 22 | TLE4 | 20591 | -7.180 | -0.5084 | Yes | ||
| 23 | SMAD4 | 20832 | -7.924 | -0.4752 | Yes | ||
| 24 | CREBBP | 21411 | -10.605 | -0.4425 | Yes | ||
| 25 | GSK3B | 21455 | -10.935 | -0.3835 | Yes | ||
| 26 | YWHAE | 21814 | -16.134 | -0.3099 | Yes | ||
| 27 | BCL9 | 21845 | -17.277 | -0.2149 | Yes | ||
| 28 | LEF1 | 21874 | -18.255 | -0.1144 | Yes | ||
| 29 | NLK | 21913 | -21.118 | 0.0016 | Yes |